NGS Library quantification products
Accurate quantitation of the number of molecules in a library is the most critical step in the preparation steps for next-generation sequencing (NGS) to obtain high-quality reading data. Indeed, in order to obtain reliable and quality reading data, it is necessary to use a sufficient quantity of DNA, and in particular for Illumina® sequencing. The use of insufficient DNA will result in low sequencing efficiency and DNA overabundance will increase cluster density and result in poor quality data.
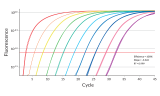
Standard methods of quantification of libraries for NGS, such as those based on electrophoresis or spectrophotometry, have low sensitivity, are not specific to the DNA bound to the adapter, and generally require a large amount of library samples for analysis. Quantitative PCR (qPCR) is considered to be the most accurate and efficient method of quantification of the library, providing consistency and reproducibility considerably higher than electrophoresis or spectrophotometry, which measure the total nucleic acid concentration. Amplification-based methods quantify only molecules that contain the two adapter sequences, thus providing a more accurate estimate of the concentration of the library molecules that can be sequenced.




